
For a full list of BASHing data blog posts see the index page. ![]()
A muggle's guide to AWK arrays: 3
Part 2 in this series looked at the 2-file command structure, where the first part of an AWK command created an array based on the first file, and the second part of the command used the array to filter a second file.
Another way to think about this command structure is that an AWK array is like a lookup table, held in memory. You can use that lookup table for different kinds of data operations on another file. In this post I'll demonstrate reformatting and table joining.
Reformatting. This example is based on the tab-separated "months" table I described in an earlier BASHing data post:
| 1 | 01 | Jan | January | i | I |
| 2 | 02 | Feb | February | ii | II |
| 3 | 03 | Mar | March | iii | III |
| 4 | 04 | Apr | April | iv | IV |
| 5 | 05 | May | May | v | V |
| 6 | 06 | Jun | June | vi | VI |
| 7 | 07 | Jul | July | vii | VII |
| 8 | 08 | Aug | August | viii | VIII |
| 9 | 09 | Sep | September | ix | IX |
| 10 | 10 | Oct | October | x | X |
| 11 | 11 | Nov | November | xi | XI |
| 12 | 12 | Dec | December | xii | XII |
For reasons I don't understand, a European organisation has recently recommended that certain scientific papers should have dates in the format d(d) Mmm. YYYY (e.g. 3 Feb. 2009) rather than in the widely accepted ISO 8601 date format (2009-02-03). I'll call this recommendation the "Eurosci" format. In what follows I'll re-format "American style" dates into "Eurosci".
The American-style dates come from a tab-separated part ("datelist") of a Wikipedia table of Japanese baseball stars and the dates of their debuts in American major league baseball:
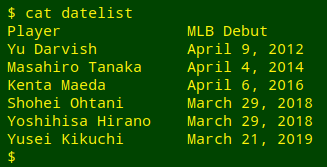
The first step in reformatting will be to break up the dates into their month, day and year pieces using AWK's split function. There are two ways to do this. The first is to use a plain space as the separator, and to remove the comma after the day string with gensub:
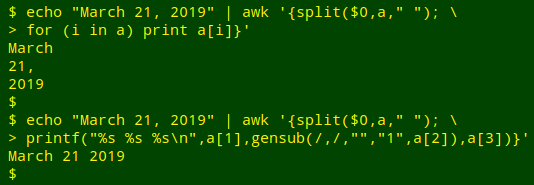
I find that method a little clumsy, so instead I'll tell split with a regex that the separator is either a space or a comma. In the demonstration case, that means the year string is the fourth element in split's array, not the third. The third element is an empty string after the comma:
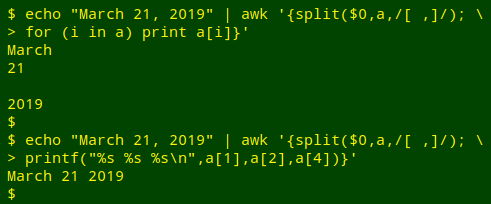
When the date string has been split into pieces, the pieces can be re-arranged, and I'll use the "months" array to replace the month string with an abbreviation:
awk -F"\t" \
'BEGIN {print "Player" FS "MLB Debut" FS "date in Eurosci"} \
FNR==NR {a[$4]=$3; next} \
FNR>1 {split($2,b,"[, ]"); \
printf("%s\t%s %s. %s\n",$0,b[2],a[b[1]],b[4])}' \
months datelist
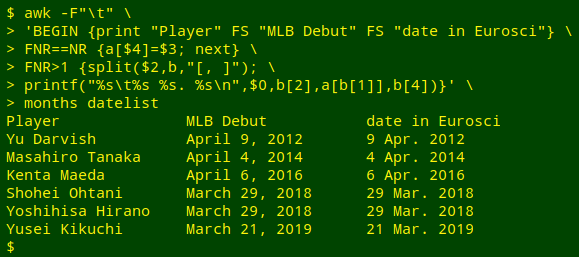
The command begins by telling AWK that the field separator in the files to be processed is a tab (-F"\t").
The BEGIN statement prints a new header for the table, using a tab as field separator (FS).
The FNR==NR condition (see part 2 of this series) directs AWK to first build an array "a" from the file "months". The index string will be the full month name, which is field 4 in "months", and the value string will be the 3-letter month abbreviation, which is field 3 (a[$4]=$3).
AWK next goes to the "datelist" file but ignores its first line (FNR>1; note the use of FNR for the current file, not NR for all lines processed so far).
The action taken on the second and succeeding lines of "datelist" begins with a split of the date string in field 2 into its components, which are stored in an array "b".
The last action in the AWK command is a printf. The first part is the formatting instruction and could be read as "[string][tab][string][space][string][full stop][space][string][newline]". The four strings to be printed are
(1) the existing line with its 2 fields ($0)
(2) the day string in "b" (b[2])
(3) the value string from the "months" array "a" where the index string is the month string in "b" (a[b[1]])
(4) the year string in "b" (b[4]).
Table joins. Before I retired as a millipede-ologist I stored specimen data in 3 plain-text, tab-separated tables:
- an events table with a unique ID number (="EventCode") for every collecting site visit
- a names table with a unique name string for each species
- a specimens table with specimen details for a name string + EventCode combination
I entered data into each table with a YAD form and did certain data ops (such as data validation and editing) independently on each table. When the time came to combine or analyse data across tables, I used a suitable AWK command based on an array or arrays. The operations were similar in their logic to SQL queries and often involved joining the tables.
For example, suppose I wanted a tab-separated list of basic facts about specimens of the millipede Lissodesmus horridomontis, together with the lat/lons and dates of collection. I could join the events and specimens tables like this:
awk -F"\t" 'FNR==NR {a[$1]=($4 FS $5 FS $15); next} \
$2 ~ /horridomontis/ {print $4 FS $5 FS $6 FS a[$3]}' events specimens \
| sort -t $'\t' -k6
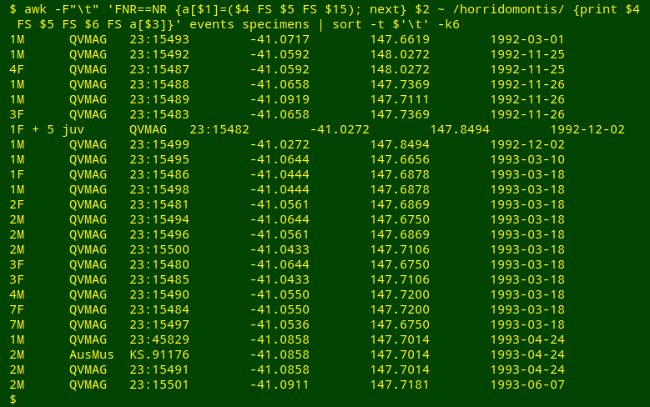
AWK first builds an array "a" from events using "EventCode" (field 1) as index string and a tab-separated combination of fields as value string: decimal latitude (field 4), decimal longitude (field 5) and ISO 8601 event date (field 15).
Moving to specimens, AWK selects all records in which name string (field 2) matches "horridomontis", and prints (with tab separation, FS) 4 fields: "Specimens" (field 4), "Repository" (field 5), "RegNo" (field 6), and the value string in "a" corresponding to that record's "EventCode" (in field 3).
Field numbers for specimens and events were always at hand in the terminal with the fields function.
Another query might be From which Australian States have millipedes in the family Haplodesmidae been collected? The AWK command will need to build arrays from events and names before processing specimens, and will therefore be joining 3 tables.
awk -F"\t" 'ARGIND==1 {a[$1]=$3; next} \
> ARGIND==2 && $6 == "Haplodesmidae" {b[$2]; next} \
> ARGIND==3 && $2 in b {print a[$3]}' \
> events names specimens | sort | uniq
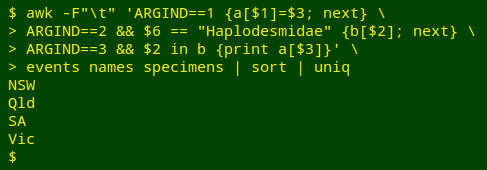
This is a 3-file AWK command that doesn't use FNR==NR to move between files. Instead it names each file to be processed with the GNU AWK variable ARGIND, which keeps track of command arguments.
AWK first builds an array "a" from events, with "EventCode" (field 1) as index string and "State" abbreviation (field 3) as value string.
AWK moves next to names. For records in which "Family" (field 6) is Haplodesmidae, AWK builds an array "b" indexed on name string (field 2).
In specimens, AWK looks for records in which name string (field 2) is an index string in "b". For those records, AWK prints the value string ("State") from array "a" for which the index string is "EventCode" (field 3).
The AWK command output is sorted and uniquified.
Last update: 2020-01-13
The blog posts on this website are licensed under a
Creative Commons Attribution-NonCommercial 4.0 International License